
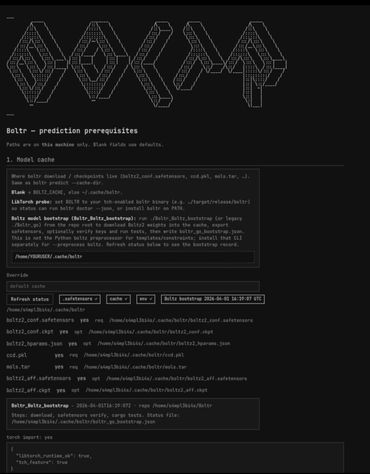
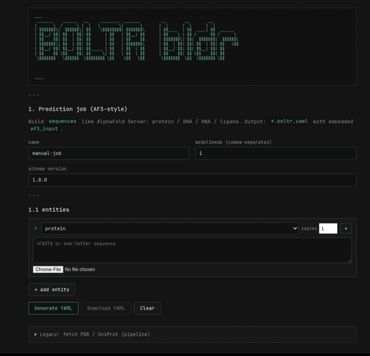
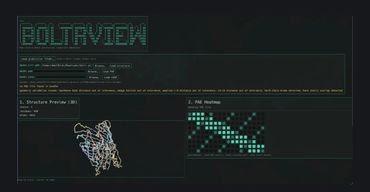
Boltr is a Rust-first protein structure prediction system designed to run Boltz/Boltz2-style workflows with stronger performance, safety, and portability than a Python-heavy stack. It takes structured protein inputs such as YAML, runs inference through a Rust + LibTorch/CUDA pipeline, and outputs files like .cif/.pdb, metadata JSON, and .a3m alignments for downstream visualization and analysis in tools like PyMOL. This program is a key component of the larger project(s) stellarOS and nucleogenesis.
// The Science of Code, Destruction and Other Synthetic Delusions of the Syndicate //
This website uses cookies.
We use cookies to analyze website traffic and optimize your website experience. By accepting our use of cookies, your data will be aggregated with all other user data.